Research News: Study Reveals Novel Therapeutic Target to Eliminate Unwanted and Misfolded Proteins
October 04, 2023
A recent discovery unearths the hidden mechanism of “aggrephagy,” with major implications for degenerative protein diseases.
In cells, the ubiquitin‒proteasome system (UPS) plays a key role in the elimination of unwanted or misfolded proteins. When UPS fails, cells activate a backup process called “aggrephagy” for clearing ubiquitin-tagged proteins. However, the associated mechanism behind this process remains unknown. Recently, a Japanese team has demonstrated how another protein called NRF1 facilitates aggrephagy, thereby providing new therapeutic targets for diseases resulting from misfolded proteins.
Biological cells contain in-built “housekeeping” mechanisms for taking care of damaged cellular structures. This includes the ubiquitin‒proteasome system (UPS), which selectively tags unwanted proteins with the ubiquitin molecule, and then clears them. When the UPS mechanism fails, cells activate a compensatory protein clearance process called “aggrephagy,” in which protein aggregates are degraded by the cell in a controlled manner. However, thus far, the mechanism behind aggrephagy has been unknown.
Now, a landmark paper published on 1 September 2023 in Volume 13, Issue 1 of Scientific Reports, reveals the underlying mechanism. “Our study demonstrates that the activation of aggrephagy is induced by the transcription factor NRF1 (NFE2L1) in response to proteasome dysfunction,” explains Atsushi Hatanaka, graduate student at Doshisha University, Japan, who is the first author of the study. The research team also included Sota Nakada and Akira Kobayashi, both from the Laboratory for Genetic Code, Graduate School of Life and Medical Sciences, Doshisha University.
NRF1, a protein involved in the transcription of DNA to RNA, plays a key role in the balance and regulation of proteins. It upregulates proteasome genes when the proteasome is damaged. As a part of the study, the team first used small interfering RNA (siRNA) to reduce the activity of the NRF1 synthesis gene in a cellular model. They then used the inhibitor MG132 to block proteasome-mediated protein recycling. These treatments led to the accumulation of undesired proteins in the cellular model, indicating that the absence of NRF1 effectively inhibited aggrephagy activation, which is otherwise typically prevalent in a cell.
The researchers then examined the effect of proteasome inhibition on genes that were direct targets of NRF1. During the course of experiments, the team noticed that in response to proteasomal failure, NRF1 caused an increase in the levels of the autophagy-related genes p62 and GABARAPL1. The elevated levels of the corresponding proteins p62 and GABARAPL1 resulted in the removal of proteins that were tagged for removal by the housekeeping protein ubiquitin. In other words, NRF1 was found to trigger the process of aggrephagy via p62 and GABARAPL1 as a direct physiological response to proteasomal failure.
Next, the team also discovered that the presence of NRF1 was necessary for the formation of p62-positive puncta—tiny round cellular structures loaded with large amounts of the protein p62. Moreover, it became evident that the colocalization (physical proximity) of the proteins p62, ULK1, and TBK1 was necessary for the activation of p62. This activation is considered to be the direct result of phosphorylation—the addition of phosphate groups to proteins. A series of experiments revealed that the phosphorylation of p62 was facilitated by NRF1. The phosphorylated p62 then contributed to the process of aggrephagy. As mentioned before, a similar increase in aggrephagy was also observed after the NRF1-mediated increase in the levels of the protein GABARAPL1.
Explaining the novelty of the research, Mr. Hatanaka says, “Although NRF1 has been previously shown to upregulate proteasome genes when the proteasome is dysfunctional, our genome-wide transcriptome analyses showed that NRF1 directly upregulates autophagy-related genes p62 and GABARAPL1.”
These findings pave the way toward the development of novel therapeutics for degenerative diseases such as Alzheimer’s disease, Parkinson’s disease, and dementia with Lewy bodies—all of which are caused by the accumulation of misfolded proteins.
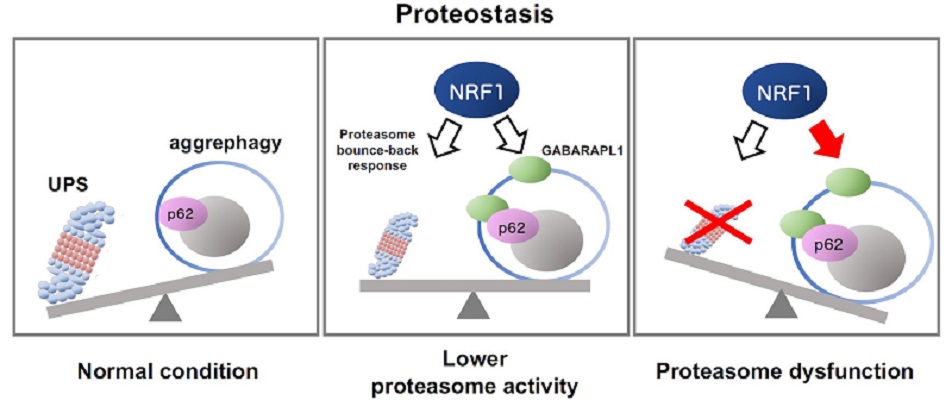
Schematic representation of NRF1-triggered activation of aggrephagy mediated by impaired proteasome activity
The ubiquitin‒proteasome system (UPS) and autophagy are protein degradation pathways essential for maintaining protein balance and regulation, or proteostasis (left panel). When proteasome activity decreases because of several reasons including chemical inhibitors and aging, the transcription factor NRF1 gets activated, leading to the upregulation of proteasome gene expression (“the proteasome bounce-back response;” middle panel). Furthermore, complete proteasome dysfunction activates NRF1-mediated aggrephagy, inducing the expression of the aggrephagy-related genes p62 and GABARAPL1 (right). These cellular responses help combat proteasome dysfunction by maintaining proteostasis.
Image courtesy: Atsushi Hatanaka, Sota Nakada, Gen Matsumoto, Katsuya Satoh, Iori Aketa, Akira Watanabe, Tomoaki Hirakawa, Tadayuki Tsujita, Tsuyoshi Waku, and Akira Kobayashi
Image license: CC BY 4.0
Image link : https://www.nature.com/articles/s41598-023-41492-9
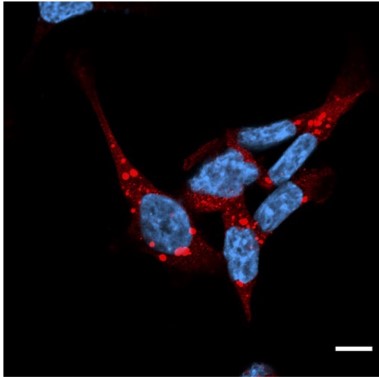
Fluorescence image of p62-positive puncta obtained using a confocal laser microscope
NRF1 plays a critical role in the formation of p62-positive puncta, facilitating the aggregation of p62 proteins after proteasome inhibition. This process is crucial for the efficient clearance of damaged proteins through aggrephagy and helps maintain cellular protein quality.
Image courtesy: Atsushi Hatanaka, Sota Nakada, Gen Matsumoto, Katsuya Satoh, Iori Aketa, Akira Watanabe, Tomoaki Hirakawa, Tadayuki Tsujita, Tsuyoshi Waku, and Akira Kobayashi
Image license: CC BY 4.0
Image link : https://www.nature.com/articles/s41598-023-41492-9
Reference
| Title of original paper | The transcription factor NRF1 (NFE2L1) activates aggrephagy by inducing p62 and GABARAPL1 after proteasome inhibition to maintain proteostasis |
| Journal | Scientific Reports |
| DOI | 10.1038/s41598-023-41492-9 |
Funding information
This work was supported in part by grants-in-aid (18J20672 (AH), 19K22826 (AK), 20H04135 (AK), 21K19743 (AK), 22H04659 (AK), 19K07650 (TW), 22K07219 (TW) and 22H03515 (TT)) from the Ministry of Education, Culture, Sports, Science and Technology and the Mitsubishi Foundation (AK).
Profiles

Atsushi Hatanaka
PhD student , Faculty of Life and Medical Sciences.
Atsushi Hatanaka is currently pursuing his Ph.D. from Doshisha University’s Graduate School of Life and Medical Sciences. He has published his research in more than seven peer-reviewed journals and his work has received more than 100 global citations so far. Atsushi has primarily conducted research in the area of molecular biology.

Akira Kobayashi
Professor , Faculty of Life and Medical Sciences Department of Medical Life Systems
Professor Akira Kobayashi received a Ph.D. degree from Tohoku University in 1997. After several years of work at the University of Tsukuba and Tohoku University, he joined the Department of Medical Life Systems, Faculty of Life and Medical Sciences, Doshisha University in 2008. He has more than 70 publications to his credit thus far. His research areas are Molecular biology and Medical biochemistry regarding cancer. Kobayashi is an association member of the Biomedical Society for Stress Response, Japan, the Japanese Cancer Association, the Japanese Biochemical Society, and the Molecular Biology Society of Japan.
Media contact
Organization for Research Initiatives & Development
Doshisha University
Kyotanabe, Kyoto 610-0394, JAPAN
CONTACT US
